Making DICOM files for neuronavigation
A user may use results of dipoles in neuronavigation system.
To do so, a user can utilize DICOM files imbedded information of dipoles.
In this page, how to make dipole-imbedded DICOM files is explained.
Beforehand, you have to confirm access authority of /neuro/setup/dicom/dicom_noes
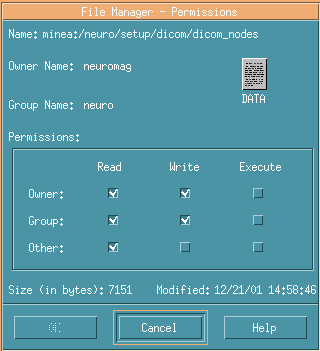
Right-click dicom_nodes, and select Change Permissions...
Confirm permission check of Write in Group.
No check in Group in the below image.
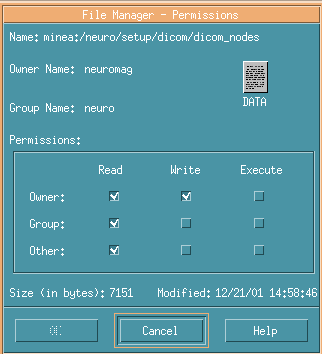
In such cases, you have to login as Owner and check Write in the Group.
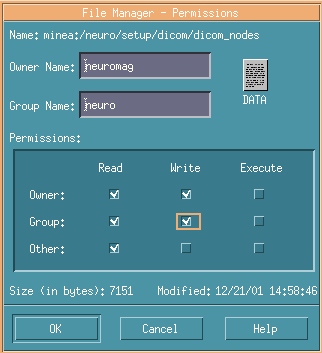
It's OK if you can see like the below image.
Then you have to copy target DICOM files.
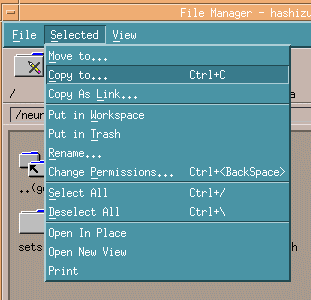
Right-click the slices including the target DICOM files and select Copy to ...
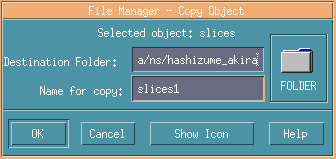
Write new names at Name for copy, here named as slice1.
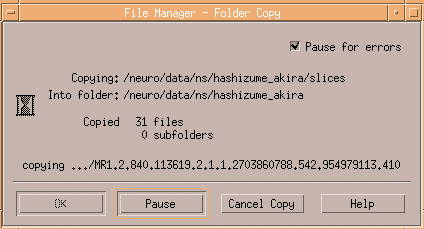
The below image shows copying DICOM files in slices to slices1.
Copying is finshed after closing the window.
Start MRIlab, and select Objects->Objects Preference.
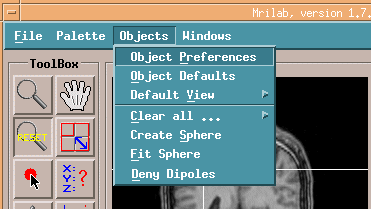
Dipole vectors shoule be None and Slice thickness shoule be 1.
Then push Apply button.
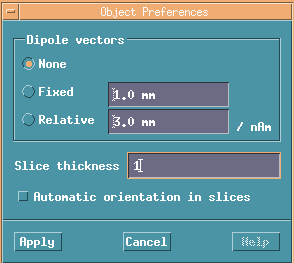
No moment lines of dipoles.
Dipoles appear on images only within 1mm from the actual dipole positions.
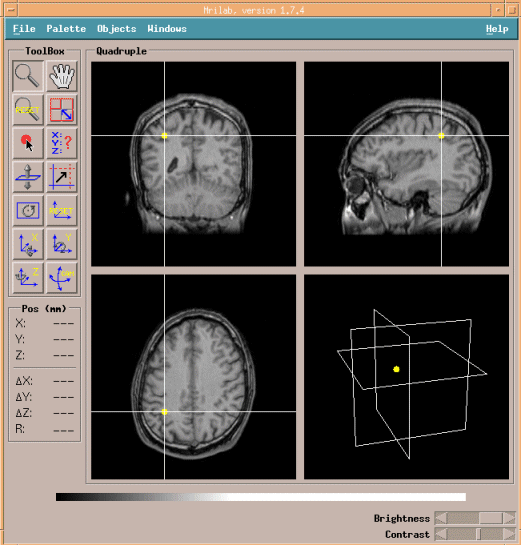
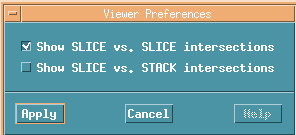
To erase white cross line, right-click on images and select Mode Prefs,
then check-off Show SLICE vs.SLICES intersections.
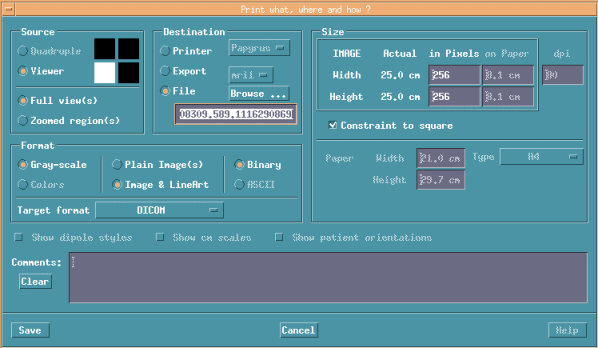
Select File->Print.

Click Viewer to select preferred cross-section images.
The below image shows horizontal cross-section images.
Select File in Destination and push Browse.. button.
Here, select the first file within the newly made folder.
Never select original slices folder, if not, you can't see MRI images on MRILAB again.
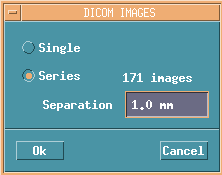
Selct Series, decide preferred slice gaps, and push OK button.
MRILAB askes where you permit OVERWITE or not.
Push Yes button.
Data is storing in some files under /var/tmp/.
Finally, new DICOM fiels are stored in folders appointed in Destination window.
You can confirm MRI1.xxx.dcm in slices1.
These are DICOM files including dipoles for neuronavigation system.
Acutually made MRI.xx.dcm files are shown.
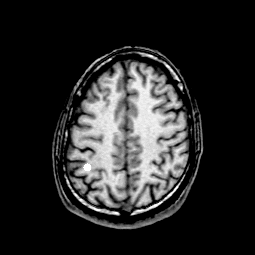
Dipoles are shown as white dots.
Where, condition of window window and window level inherited to the image condision of MRILAB.



















